New model allows pharmacological researchers to dock nearly any drug and see how it behaves in P-glycoprotein, a protein in the cell associated with failure of chemotherapy
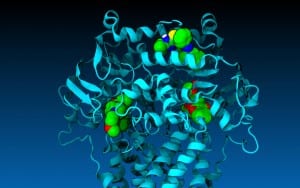
Drugs important in the battle against cancer responded the way they do in real life and behaved according to predictions when tested in a computer-generated model of one of the cell’s key molecular pumps — the protein P-glycoprotein, or P-gp.
Biologists at Southern Methodist University, Dallas, developed the computer generated model to overcome the problem of relying on only static images for the structure of P-gp, said biologist John G. Wise, lead author on the journal article announcing the advancement.
The new SMU model allows researchers to dock nearly any drug in the P-gp protein and see how it will actually behave in P-gp’s pump, said Wise, an associate professor in SMU’s Department of Biological Sciences.
“The value of this fundamental research is that it generates dynamic mechanisms that let us understand something in biochemistry, in biology,” he said. “And by understanding P-gp in such detail, we can now think of ways to better and more specifically inhibit it.”
P-gp is the cellular pump that protects cells by pumping out toxins. But that’s a problem when P-gp targets chemotherapy drugs as toxic, preventing chemo from killing cancer cells. Scientists are searching for ways to inhibit P-gp’s pumping action.
The SMU researchers tested Tariquidar, a new P-gp inhibitor still in clinical trials. Inhibitors offer hope for stopping P-gp’s rejection of chemotherapeutics by stalling the protein’s pumping action. Pharmacology researchers disagree, however, on where exactly Tariquidar binds in P-gp.
When run through the SMU model, Tariquidar behaved as expected: It wasn’t effectively pumped from the cell and the researchers observed that it prefers to bind high in the protein.
“Now we have more details on how Tariquidar inhibits P-gp, where it inhibits and what it’s actually binding to,” Wise said.

Also using the model, the researchers discovered greater detail than previously known about the behavior of other drugs as well, and how those drugs bind in P-gp to stop its pumping action.
The study was funded in part by the National Institutes of Health. The lab was recently awarded a second NIH grant for the research.
The findings are published in the journal Biochemistry. The article, “Multiple drug transport pathways through human P-glycoprotein,” is published online in advance of print at NIH’s PubMed Central.
A still image of the modeled protein in action will appear on the cover of the October through December issues of Biochemistry.
Testing the virtual P-gp model by virtually docking real drugs
Wise and his colleagues tested one of the workhorse drugs of chemotherapy, daunorubicin, a close cousin of Adriamycin.
An aggressive chemotherapeutic, daunorubicin stops DNA replication in the cell, and is a classic target for P-gp to pump out of a cell, Wise said.
“For a long time, it’s been thought that there are at least a couple of distinct binding sites for drugs,” Wise said. “Sure enough, with our models, we found that daunorubicin, at least, prefers to bind on one side of the P-gp model, while verapamil – a commonly prescribed blood pressure medicine – prefers the other side.”

Not only did the researchers show computationally that there are two different starting points for drugs, they also showed that there are two different pathways to get the drugs through.
“The two different drugs start at different sites and they’re funneled to the outside by being pushed by the protein,” Wise said. “But the actual parts of the protein that are pushing the drugs out are different.”
Wise and his co-authors, SMU biologists Pia Vogel and James McCormick, created the P-gp computer-generated simulation using SMU’s High Performance Computer, ManeFrame.
Molecular model can aid in fight against multi-drug resistance of cancer cells
The capability of watching molecular machinery up close, while doing its job the way it does in real life, may spark new drug discoveries to fight cancer.
“Having an accurate model that actually moves – that shows the dynamics of the thing – is incredibly helpful in developing therapies against a molecular target to inhibit it. The only other ways to do it are blind, and the chances of success using blind methods are very low,” Wise said.
“Scientists have tried for 30 years to find inhibitors of this pump and have done it without knowing the structure and with only little knowledge about the mechanism, screening more or less blindly for compounds that inhibit the thing,” Wise said. “They found drugs that worked in the test tube and that worked in cultured cells, but that didn’t work in the patient. With our model, because we can see the pump moving, we can probably predict better what’s going to make an inhibitor actually work well.”
Vogel and Wise led a team of researchers in using the P-gp model to virtually screen millions of publically available drug-like compounds.

They discovered three new drug leads that could ultimately inhibit P-gp and offer better odds of survival to prostate cancer patients. The researchers reported those findings this month in the journal Pharmacology Research & Perspectives, http://bit.ly/1XGjN5w.
New SMU model simulates molecular machinery in action
Researchers look for drug compounds that can temporarily stop or inhibit the P-gp pump, so that the chemotherapy drugs that enter the cancer cell will stay there and do the job of killing the cancer. Finding the right pump inhibitor requires understanding the pumping action. That’s difficult without seeing the pump at work.
The structures of proteins similar to P-gp have been previously available in a static state through X-ray crystallography. Scientists use X-ray crystallography as a tool that essentially draws the details of biological structures by identifying their atomic and molecular structure through diffraction of X-rays by the atoms themselves.
Scientists often contribute the resulting protein structures to the U.S. Protein Data Bank repository for public use.
Detailed data combined with several trillion calculations produced model
To build the P-gp model, Wise used structures from the repository, showing various stages of transport, to simulate four points of reference. From there, SMU’s ManeFrame supercomputer was fed parameters and characteristics of the protein as well as how it should behave physically, including when kinetic energy was added to bring the protein and its surrounding membrane and water up to body temperature. The animated model resulted from calculating differences between two structures and using targeted molecular dynamics programs to slightly nudge the model to the next step.
“You do that several million times and make several trillion calculations and you arrive at the next structure,” Wise said. “In this way, we can nudge P-gp through a full catalytic transport cycle.”
Finally, using a docking program, the researchers individually introduced daunorubicin and other drugs into the protein, and watched the drugs move through P-gp’s catalytic cycle.
“What happened was — the drugs moved,” Wise said. “And they moved the way they should move, clinically, biochemically, physiologically, to pump the compounds out of the cell.”
Vogel added that, “in some of the zoom-ins of the model, you can actually see the amino acids paddle down the drugs.”
Further challenging and testing the model
The researchers ran a critical control to further test if the model worked.
“We thought maybe anything you put in the protein, relevant or not, would get pumped through. So we put in something that is not a transport substrate of P-gp, something that biochemically would never be transported by P-gp,” Wise said. “We put it in, starting where daunorubicin is effectively pumped out, and very quickly the compound left the protein — but it left the opposite way, back into the cell. This experiment gave us more confidence that what we are seeing in these models is reflecting what happens in the cell.”
Wise admits that until he saw it for himself, even he had doubts the virtual P-gp model would behave like real-life P-gp.
“It’s a crude approximation of a complex, sophisticated human protein, but it’s so much better than the static images available now,” Wise said. “I’ve got to emphasize for all the disbelievers, for the ‘culture of doubters’ out there, that this model works — it moves the drugs through the membrane. That speaks for itself. What P-gp does in the cell, cancerous or normal, it does in our simulations.”









 Department of Defense awards $2.6 million to SMU STEM program for minority students
Department of Defense awards $2.6 million to SMU STEM program for minority students
 Women have made strides for equality in society, but gender gap still exists in art museum directorships
Women have made strides for equality in society, but gender gap still exists in art museum directorships Search for dark matter covers new ground with CDMS experiment in Minnesota
Search for dark matter covers new ground with CDMS experiment in Minnesota NOvA experiment glimpses neutrinos, one of nature’s most abundant, and elusive particles
NOvA experiment glimpses neutrinos, one of nature’s most abundant, and elusive particles SMU scientists to deploy seismic monitors in North Texas region near Azle, Texas
SMU scientists to deploy seismic monitors in North Texas region near Azle, Texas Observed by Texas telescope: Light from huge explosion 12 billion years ago reaches Earth
Observed by Texas telescope: Light from huge explosion 12 billion years ago reaches Earth Low IQ students learn to read at 1st-grade level after persistent, intensive instruction
Low IQ students learn to read at 1st-grade level after persistent, intensive instruction Richest marine reptile fossil bed along Africa’s South Atlantic coast is dated at 71.5 mya
Richest marine reptile fossil bed along Africa’s South Atlantic coast is dated at 71.5 mya


 Observed! SMU’s LHC physicists confirm new particle; Higgs ‘God particle’ opens new frontier of exploration
Observed! SMU’s LHC physicists confirm new particle; Higgs ‘God particle’ opens new frontier of exploration DOE Award: advancing SMU’s link to the God particle
DOE Award: advancing SMU’s link to the God particle
 Ancient tree-ring records from southwest U.S. suggest today’s megafires are truly unusual
Ancient tree-ring records from southwest U.S. suggest today’s megafires are truly unusual Human diabetes has new research tool: Overfed fruit flies that develop insulin resistance
Human diabetes has new research tool: Overfed fruit flies that develop insulin resistance